Shen
Laboratory
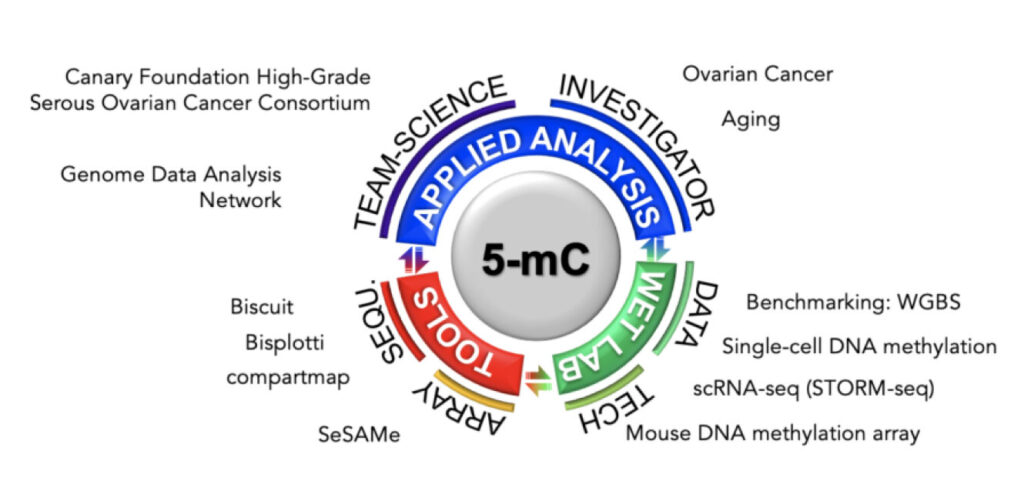 Epigenomic Analysis in Human Disease
Epigenomic Analysis in Human Disease
The Shen Laboratory focuses on the epigenome and its interaction with the genome in various diseases, with a specific emphasis on female cancers and cross-cancer comparisons. The Shen team uses bioinformatics as a tool to understand the etiology, cell of origin, and epigenetic mechanisms of various diseases and to devise better approaches for cancer prevention, detection, therapy, and monitoring. Team members have extensive experience with genome-scale DNA methylation profiles in primary human samples and have made major contributions to epigenetic analysis within The Cancer Genome Atlas (TCGA).
DNA methylation is ideally suited to deconstruct heterogeneity among cell types within a tissue sample. In cancer research, this approach can be used for both cancer cell clonal evolution studies or for quantifying normal cell infiltration and stromal composition. The latter can provide insights into the tumor microenvironment. For non-cancer studies, this method is a useful tool to accurately estimate different cell populations and provide insights into lineage structures and population shifts in disease. The Shen lab also is interested in translational applications of epigenomic technology. To this end, the lab translates markers that emerge from bioinformatics analysis into clinical assay development, marker panel assembly, and optimization, with the ultimate goal of clinical testing and validation.
Featured News & Featured Publications
Learn More
New ‘atlas’ may aid search for markers of early-stage ovarian cancer
Van Andel Institute scientists develop technique for high-resolution single cell epigenetic analysis

Van Andel Institute’s Dr. Hui Shen elected to the AIMBE College of Fellows
Dror E… Shen H, Pospisilik JA. 2023. Epigenetic dosage identifies two major and functionally distinct β cell subtypes. Cell Metab.
Brown JR … Shen H … Buckanovich RJ. Phase II clinical trial of metformin as a cancer stem cell–targeting agent in ovarian cancer. JCI Insight.
Carrot-Zhang J .. Shen H … Ziv E. Comprehensive analysis of genetic ancestry and its molecular correlates in cancer. Can Cell.
Gao GF … Shen H … Noble MS. Before and after: Comparison of legacy and harmonized TCGA Genomic Data Commons’ data. Cell Sys.
Wilson MR … Shen H … Chandler RL. ARID1A and PI3-kinase pathway mutations in the endometrium drive epithelial transdifferentiation and collective invasion. Nat Commun.
Our Impact
We’re raising thousands to save millions.
We’re turning hope into action for the millions of people around the world affected by diseases like cancer and Parkinson’s. Find out how you can help us make a difference.
- 141 peer-reviewed papers published in 2025, 74 of which were in high-impact journals
- 15 VAI-SU2C Epigenetics Dream Team clinical trials launched to date
- 10 clinical trials and related projects supported by VAI through the International Linked Clinical Trials Program
Hui Shen, Ph.D.
Professor, Department of Epigenetics
Areas of Expertise
Epigenomics, DNA methylation, bioinformatics, ovarian cancer, pan-cancer comparison, genome-epigenome interaction
Biography
An expert in bioinformatics and computational biology, Dr. Hui Shen develops and applies cutting-edge genomics technologies and bioinformatic tools to investigate the regulatory mechanisms of gene transcription and cell differentiation — collectively known as epigenetics. Her multi-faceted research has led to improved understanding of molecular and cellular mechanisms driving various cancer types, particularly ovarian cancer.
Dr. Shen earned her B.Sc. in biology from Nanjing University and her Ph.D. in genetic, molecular and cellular biology from the University of Southern California (USC), where she was supported by a Provost Ph.D. Fellowship. She was part of The Cancer Genome Atlas (TCGA) team, a landmark multi-institutional effort to map the molecular basis of cancer. Dr. Shen joined Van Andel Institute in 2014 as an assistant professor, rising to associate professor in 2018 and full professor in 2023.
She is an F1000 Faculty Member in Cancer Epigenetics and has received numerous honors, including the Liz Tilberis Award from the Ovarian Cancer Research Alliance, an NIH/NCI ESI MERIT Award, and an AACR Team Science Award. In 2025, she was elected to the College of Fellows of the American Institute for Medical and Biological Engineering (AIMBE) in recognition of her contributions to computational cancer epigenetics.
Selected Publications
For a full list of Dr. Shen’s publications, please visit Google Scholar.
2026
Burkhard KM, Semwal A, Johnson BK, Chu KC, Kranick RJ, Rayan M, DiFeo A, Shen H, Mehta G. Preprint. Tumoroid model recreates clinically relevant phenotypes of high grade serous ovarian cancer cells, carcinoma associated fibroblasts, and macrophages. Res Sq.
2025
Oswald BM, DeCamp LM, Longo J, Dahabieh MS, Bunda N, Johnson BK, Watson MJ, Ma S, Preston SEJ, Sheldon R, Vincent MP, Ellis AE, Sopper-Hopper MT, Isaguirre C, Kamarudin D, Shen H, Williams KS, Crawford PA, Kaech S, Jang HJ, Lien EC, Krawczyk CM, Jones RG. 2025. Dietary restriction reprograms CD8+ T cell fate to enhance anti-tumor immunity and immunotherapy responses. Nat Metab.
Spix NJ, Habib WA, Zhang Z, Eugster E, Milliron HY, Sokol D, Lee K, Nolte PA, Endicott JL, Krzyzanowski KF, Hinoue T, Morrison J, Johnson BK, Zhou W, Shen H, Laird PW. 2025. High-coverage allele-resolved single-cell DNA methylation profiling reveals cell lineage, X-inactivation state, and replication dynamics. Nat Commun 16(1):6273.
Bhinder B, Friedl V, Sethuraman S, Risso D, Chiotti KE, Mashl RJ, Ellrott KP, Lee JA, Wong CK, Gyan K, Deshpande A, Imielinski M, Bareja R, Stuart J, Peto M, Hoadley KA, Lazar AJ, Cherniack AD, Zhu J, Cao S, Rubin M, Wang W, Bathe OF, Robine N, Ding L, Laird PW, Zhou W, Shen H, Thorsson V, Yeh JJ, Bailey MH, Zhou DC, Peng XL, Goldman M, Li Y, Korkut A, Sahni N, Hayes DN, Mensah MKA, Felau I, Kemal A, Caesar-Johnson S, Demchok JA, Yang L, Ferguson ML, Tarnuzzer R, Wang Z, Zenklusen JC, Cancer Genome Atlas Analysis Network, Spellman P, Elemento O. 2025. Pan-cancer immune and stromal deconvolution predicts clinical outcomes and mutation profiles. Sci Rep 15(1):23921.
Beddows I, Djirackor S, Omran DK, Jung E, Shih NNC, Roy R, Hechmer A, Ohshen A, Adelmant G, Tom A, Morrison J, Adams M, Rohrer DC, Schwartz LE, Pearce CL, Auman H, Marto JA, Drescher CQ*, Drapkin R*, Shen H*. 2025. Impact of BRCA mutations, age, surgical indication, and hormone status on the molecular phenotype of the human Fallopian tube. Nat Commun 16:2981.
*Co-corresponding authors
2024
Kaymak I, Watson MJ, Oswald BM, Ma S, Johnson BK, DeCamp LM, Mabvakure BM, Luda KM, Ma EH, Lau KH, Fu Z, Muhire B, Kitchen-Goosen SM, VanderArk A, Dahabieh MS, Samborska B, Vos M, Shen H, Fan ZP, Roddy TP, Kingsbury GA, Sousa CM, Krawczyk CM, Williams KS, Sheldon RD, Kaech SM, Roy DG, Jones R. 2024. ACLY and ACSS2 link nutrient-dependent chromatin accessibility to regulation of CD8 T cell effector responses. J Exp Med 221(9).
Spix NJ, Abi Habib W, Zhang Z, Eugster E, Milliron H, Sokol D, Lee K, Nolte P, Endicott J, Krzyzanowski KF, Hinoue T, Morrison J, Johnson BK, Zhou W, Shen H, Laird PW. Pre-print. High-coverage allele-resolved single-cell DNA methylation profiling by scDEEP-mC reveals cell lineage, X-inactivation state, and replication dynamics. bioRxiv.
Zhou W, Johnson BK, Morrison J, Beddows I, Eapen J, Katsman E, Semwal A, Habib W, Heo L, Laird PW, Berman B, Triche Jr TJ, Shen H. 2024. BISCUIT: an efficient, standards-compliant tool suite for simultaneous genetic and epigenetic inference in bulk and single-cell studies. Nuc Acids Res 52(6):e32.
Beddows I, Fan H, Heinze K, Johnson BK, Leonova A, Senz J, Djirackor S, Cho KR, Pearce CL, Huntsman DG, Angelesio MS, Shen H. 2024. Cell state of origin impacts development of distinct endometriosis-related ovarian carcinoma histotypes. Can Res 84(1):26–38.
2023
Morrison J, Zhou W, Johnson BK, Shen H. 2023. Dupsifter: A lightweight duplicate marking tool for whole genome bisulfite sequencing. Bioinformatics 39(12):btad729.
*Liang W-W, Lu RJ-H … Shen H … Wyczalkowksi MA. 2023. Integrative multi-omic cancer profiling reveals DNA methylation patterns associated with therapeutic vulnerability and cell-of-origin. Can Cell 41(9):1567–1585.
Caramia F, Speed TP, Shen H, Haupt Y, Haupt S. 2023. Establishing the link between X-chromosome aberrations and TP53 status, with breast cancer patient outcomes. Cells 12(18):2245.
Jang HJ, Hostetter G, MacFarlane AW, Madaj Z, Ross EA, Hinoue T, Kulchycki JR, Burgos RS, Tafseer M, Alpaugh RK, Schwebel CL, Kokate R, Geynisman DM, Zibelman MR, Ghatalia P, Nichols PW, Chung W, Madzo J, Hahn NM, Quinn DI, Issa JPJ, Topper MJ, Baylin SB, Shen H, Campbell KS, Jones PA, Plimack ER. 2023. A phase II trial of guadecitabine plus atezolizumab in metastatic urothelial carcinoma progressing after initial immune checkpoint inhibitor therapy. Clin Cancer Res: OF1–OF4.
*Shih J, Sarmashghi S, Zhavula-Kostadinova N, Zhang S, Georgis Y, Hoyt SH, Cuoco MS, Gao GF, Sputt LF, Berger AC, Ha G, Rendo V, Shen H, Meyerson M, Cherniack AD, Taylor AM, Beroukhim R. 2023. Cancer aneuploidies are shaped primarily by effects on tumour fitness. Nature 619(7971):793–800.
Dror E, Fagnocchi L, Wegert V, Apostle S, Grimaldi B, Gruber T, Panzeri I, Heyne S, Höffler KD, Kreiner V, Ching R, Lu TTH, Semwal A, Johnson B, Senapati P, Lempradl A, Schones D, Imhof A, Shen H, Pospisilik JA. 2023. Epigenetic dosage identifies two major and functionally distinct β cell subtypes. Cell Metab.
*Garcia-Recio S, Hinoue T … Shen H …AURORA US Network, Perou CM. 2023. Multiomics in primary and metastatic breast tumors from the aurora us network finds microenvironment and epigenetic drivers of metastasis. Nat Cancer 4(1):128–147.
2022
Garcia-Recio S, Hinoue T, Wheeler GL, Kelly BJ, Garrido-Castro AC, Pascual T, De Cubas AA, Xia Y, Felsheim BM, McClure MB, Rajkovic A, Karaesmen E, Smith MA, Fan C, Gonzalez Ericsson PI, Sanders ME, Creighton CJ, Bowen J, Leraas K, Burns RT, Coppens S, Wheless, Rezk S, Garrett AL, Parker JS, Foy KK, Shen H, Park BH, Krop I, Anders C, Gastier-Foster J, Rimawi MF, Nanda R, Lin NU, Isaacs C, Marcom PK, Stornioo AM, Couch FJ, Chandran U, Davis M, Silverstein J, Ropelewski A, Liu AC, Hilsenbeck SG, Norton L, Richardson AL, Symmans WF, Wolff AC, Davidson NE, Carey LA, Lee AV*, Balko JM*, Hoadley KA*, Laird PW*, Mardis AR*, King TA*, AURORA US Network, Perou CM*. 2022. Multiomics in primary and metastatic breast tumors from the AURORA US network finds microenvironment and epigenetic drivers of metastasis. Nat Can.
* These authors jointly supervised this work
Endicott JL, Nolte PA, Shen H, Laird PW. 2022. Cell division drives DNA methylation loss in late-replicating domains in primary human cells. Nat Comm 13:6659.
Chambwe N, Sayaman RW, Hu D, Huntsman, The Cancer Genome Atlas Network*, Kemal A, Caesar-Johnson S, Zenklusen JC, Ziv E, Beroukim R, Cherniack AD. 2022. Analysis of germline-driven ancestry-associated gene expression in cancers. STAR Protoc 3(3).
*Dr. Shen is a member of The Cancer Genome Atlas Network
Paul EN, Grey JA, Carpenter TJ, Madaj ZB, Lau KH, Givan SA, Burns GW, Chandler RL, Wegienka GR, Shen H, Teixeira JM. 2022. Transcriptome and DNA methylome analyses reveal underlying mechanisms for the racial disparity in uterine fibroids. JCI Insight 7(20): e160274.
Wiseman AK, Tiedemann RL, Fan H, Shen H, Madaj Z, McCabe MT, Pappalardi MB, Jones PA. 2022. Chromosome-specific retention of cancer-associated DNA hypermethylation following pharmacological inhibition of DNMTi. Commun Biol 5:528.
Zhou W, Hinoue T, Barnes B, Mitchell O, Iqbal W, Lee SM, Foy KK, Lee KH, Moyer EJ, VanderArk A, Koeman JM, Ding W, Kalkat M, Spix NJ, Eagleson B, Pospisilik JA, Szabó PE, Bartolomei M, Vander Scaaf NA, Kang L, Wiseman AK, Jones PA, Krawczyk CM, Adams M, Porecha R, Chen BH, Shen H, Laird PW. 2022. DNA methylation dynamics and dysregulation delineated by high-throughput profiling in the mouse. Cell Genom 2(7):100144.
Guak H, Sheldon RD, Beddows I, Vander Ark A, Weiland MJ, Shen H, Jones RG, St-Pierre J, Ma EH, Krawczyk CM. 2022. PGC-1B maintains mitochondrial metabolism and restrains inflammatory gene expression. Sci Rep 12(1): 16028.
2021
Carrot-Zhang J, Han S, Zhou W, Damrauer JS, Kemal A, The Cancer Genome Atlas Network, Cherniack AD, Beroukhim R. 2021. Analytical protocol to identify local ancestry-associated molecular features in cancer. STAR Protoc 4(17):100766.
*Dr. Shen is a member of The Cancer Genome Atlas Network
Carrot-Zhang J … The Cancer Genome Atlas Research Network … Meyerson M, Govindan R, Imielinski M. 2021. Whole-genome characterization of lung adenocarcinomas lacking alterations in the RTK/TAS/RAF pathway. Cell Rep 34(5):108707.
*Dr. Shen is a member of The Cancer Genome Atlas Network
Conley BA … Shen H … Ivy SP. 2021. The Exceptional Responders Initiative: Feasibility of a National Cancer Institute pilot study. J Natl Cancer Instit 113(1):27 37.
Morrison J, Koeman JM, Johnson BK, Foy KK, Beddows I, Zhou W, Chesla DW, Rossell LL, Siegwald EJ, Adams M, Shen H. 2021. Evaluation of whole-genome DNA methylation sequencing library preparation protocols. Epigenetics Chromatin 14:28.
Wheeler DA, Takebe N, Hinoue T, Hoadley KA, Cardenas MF, Hamilton AM, Laird PW, Wang L, Johnson A, Dewal N, Miller V, Piñeyro D, Castro de Moura M, Esteller M, Shen H, Zenklusen JC, Tarnuzzer R, McShane LM, Tricoli JV, Williams PM, Lubensky I, O’Sullivan-Coyne G, Kohn EC, Little RF, White J, Malik S, Harris L, Weil C, Chen AP, Karlovich C, Rodgers B, Shankar L, Jacobs P, Nolan T, Hu J, Muzny DM, Doddapaneni H, Korchina V, Gastier-Foster J, Bowen J, Leraas K, Edmondson EF, Doroshow JH, Conley BA, Ivy SP, Staudt LM. 2021. Molecular features of cancers exhibiting exceptional responses to treatment. Can Cell 39(1):38–53.e7.
2020
Brieger KK, Peterson S, Lee AW, Mukherjee B, Bakulski KM, Alimujiang A, Anton-Culver H, Anglesio MS, Bandera EV, Berchuck A, Bowtell DDL, Chenevix-Trench G, Cho KR, Cramer DW, DeFazio A, Doherty JA, Fortner RT, Garsed DW, Gayther SA, Gentry-Maharaj A, Goode EL, Goodman MT, Harris HR, Høgdall E, Huntsman DG, Shen H, Jensen A, Johnatty SE, Jordan SJ, Kjaer SK, Kupryjanczyk J, Lambrechts D, McLean K, Menon U, Modugno F, Moysich K, Ness R, Ramus SJ, Richardson J, Risch H, Rossing MA, Trabert B, Wentzensen N, Ziogas A, Terry KL, Wu AH, Hanley GE, Pharoah P, Webb PM, Pike MC, Pearce CL. 2020. Menopausal hormone therapy prior to the diagnosis of ovarian cancer is associated with improved survival. Gynecol Oncol 158(3):702–709.
Conley BA, Staudt L, Takebe N, Wheeler DA, Wang L, Cardenas MF, Korchina V, Zenklusen JC, McShane LM, Tricoli JV, Williams PM, Lubensky I, O’Sullivan-Coyne G, Kohn E, Little RF, White J, Malik S, Harris LN, Mann B, Weil C, Tarnuzzer R, Karlovich C, Rodgers B, Shankar L, Jacobs PM, Nolan T, Berryman SM, Gastier-Foster J, Bowen J, Leraas K, Shen H, Laird PW, Esteller M, Miller V, Johnson A, Edmondson EF, Giordano TJ, Kim B, Ivy SP. 2020. The Exceptional Responders Initiative: Feasibility of A National Cancer Institute pilot study. J Natl Cancer Inst 113(1):27–37.
The ICGC/TCGA Pan-Cancer Analysis of Whole Genomes Consortium. 2020. Pan-cancer analysis of whole genomes. Nature 578(7793):82–93.
*Shen H is a coauthor through the ICGC consortium
Fan H, Atiya HI, Wang Y, Pisanic TR, Wang TH, Shih IM, Foy KK, Frisbie L, Buckanovich RJ, Chomiak AA, Tiedemann RL, Rothbart SB, Chandler C, Shen H, Coffman LG. 2020. Epigenomic reprogramming toward mesenchymal-epithelial transition in ovarian-cancer-associated mesenchymal stem cells drives metastasis. Cell Rep 33(10):108473.
Brown JR, Chan DK, Shank JJ, Griffith KA, Fan H, Szulawski R, Yang K, Reynolds RK, Johnston C, McLean K, Uppal S, Liu JR, Cabrera L, Taylor SE, Orr BC, Modugno F, Mehta P, Bregenzer M, Mehta G, Shen H, Coffman LG, Buckanovich RJ. 2020. Phase II clinical trial of metformin as a cancer stem cell–targeting agent in ovarian cancer. JCI Insight.
Carrot-Zhang J, Chambwe N, Damrauer JS, Knijnenburg TA, Robertson AG, Yau C, Zhou W, Berger AC, Huang K, Newberg JY, Mashl RJ, Romanel A, Sayaman RW, Demichelis F, Felau I, Frampton GM, Han S, Hoadley KA, Kemal A, Laird PW, Lazar AJ, Le X, Oak N, Shen H, Wong CK, Zenklusen JC, Ziv E. 2020. Comprehensive analysis of genetic ancestry and its molecular correlates in cancer. Can Cell 5(11):639–654.
2019
Gao GF, Parker, JS, Reynolds SM, Silva TC, Wang LB, Zhou W, Akbani R, Bailey M, Bale S, Berman BP, Brooks D, Chen H, Cherniak AD, Demchok JA, Ding J, Felau I, Gaheen S, Gergard DS, Heiman DI, Hernandez KM, Hoadley KA, Jayasinghe R, Kemal A, Knijnenburg TA, Laird PW, Menash MKA, Mungall AJ, Robertson AG, Shen H, Tarnuzzer R, Wang Z, Wyczalkowski M, Yang L, Zenklusen JC, Zhang Z, The Genomic Data Analysis Network, Liang H, Noble MS. 2019. Before and after: Comparison of legacy and harmonized TCGA Genomic Data Commons’ data. Cell Sys 9(1):24–34.
Wilson MR, Reske JJ, Holladay J, Wilber GE, Rhodes M, Koeman J, Adams M, Johnson B, Su RW, Joshi NR, Patterson AL, Shen H, Leach RE, Teixeria JM, Fazleabas AT, Chandler RL. 2019. ARID1A and PI3-kinase pathway mutations in the endometrium drive epithelial transdifferentiation and collective invasion. Nat Commun 10:3554.
George JW+, Fan H+, Johnson B, Carpenter TJ, Foy KK, Chatterjee A, Patterson AL, Koeman J, Adams M, Madaj ZB, Chesla D, Marsh EE, Triche TJ, Shen H*, Teixeira JM*. 2019. Integrated epigenome, exome, and transcriptome analyses reveal molecular subtypes and homeotic transformation in uterine fibroids. Cell Rep.
+ co-first authors; * co-senior authors
2018
Zhou W, Triche Jr. TJ, Laird PW, Shen H*. 2018. SeSAMe: reducing artifactual detection of DNA methylation by Infinium BeadChips in genomic deletions. Nuc Acids Res 46(20):e123-e123.
*senior author
Shen H+, Shih J, Hollern DP, Wang L, Bowlby R, Tickoo SK, Thorsson V … Hoadley KA. 2018. Integrated molecular characterization of testicular germ cell tumors. Cell Rep 23(11):3392–3406.
+ co-first authors
Sanchez-Vega F, Mina M, Armenia J…Zhou W, Shen H, Laird PW…Schultz N. 2018. Oncogenic signaling pathways in The Cancer Genome Atlas. Cell 173(2):321–337.
*One of three theme papers for the PanCancer Atlas
Berger AC, Korkut A, Kanchi RS…Fan H, Shen H…Akbani R. 2018. A comprehensive pan-cancer molecular study of gynecological and breast cancers. Can Cell 33(4):690–705.
Knijnenburg T, Wang L, Zimmerman MT, Chambwe N…Fan H, Shen H…Laird PW…Wang C. 2018. Genomic and molecular landscape of DNA damage repair deficiency across The Cancer Genome Atlas. Cell Rep 23(1):239–254.
Campbell JD, Yau C, Bowlby R, Liu Y, Brennan K, Fan H, Taylor AM, Wang C, Walter V, Akbani R, Byers LA, Creighton CJ, Coarfa C…Shen H…Laird PW…Waes CV. 2018. Genomic, pathway network, and immunologic features distinguishing squamous carcinomas. Cell Rep 23(1):194–212.
Ricketts CJ…Fan H…Shen H…Laird PW…Spellman P, Rathmell K, Linehan WM. 2018. Comprehensive molecular characterization of renal cell carcinoma. Cell Rep 23(1):313–326.
Thorsson V…Zhou W, Shen H…Shmulevich I. 2018. The immune landscape of cancer. Immunity 48(4):812–830.
Zhou W+, Dinh HQ+, Ramjan Z, Weisenberger DJ, Nicolet CM, Shen H*, Laird PW*, Berman BP*. 2018. DNA methylation loss in late-replicating domains is linked to mitotic cell division. Nat Gen 50:591–602.
+ co-first authors; * co-senior authors
2017
Zhou W, Laird PW, Shen H. 2017. Comprehensive characterization, annotation and innovative use of Infinium DNA methylation BeadChip probes. Nuc Acids Res 45(4):e22–e22.
The Cancer Genome Atlas Research Network. 2017. Comprehensive and integrative genomic characterization of hepatocellular carcinoma. Cell. 169(7):1327–1341.
Cherniack AD, Shen H…Laird PW…The Cancer Genome Atlas Research Network..Akbani R, Levine DA. 2017. Integrated molecular characterization of uterine carcinoma. Can Cell 31(3):411–423.
2016
The Cancer Genome Atlas Research Network. 2016. Comprehensive molecular characterization of papillary renal-cell carcinoma. New Eng J Med 374(2):135–145.
Gross AM, Jaeger PA, Kreisberg JF, Loncon K, Jepsen KL, Khosroheidari M, Morsey BM, Swindells S, Shen H, Ng CT, Flagg K, Chen D, Zhang K, Fox HS, Ideker T. 2016. Methylome-side analysis of chronic HIV infection reveals five-year increase in biological age and epigenetic targeting of HLA. Mol Cell 62(2):157–168.
Li B, Severson E, Pignon JC, Zhao H, Li T, Novak J, Jiang P, Shen H, Aster JC, Rodig S, Signoretti S, Liu JS, Liu XS. 2016. Comprehensive analyses of tumor immunity: implications for cancer immunotherapy. Genome Biol 17:174.
Liu M, Ohtani H, Zhou W, Ørskov AD, Charlet J, Zhang YW, Shen H, Baylin SB, Liang G, Grønbaek K, Jones PA. 2016. Vitamin C increases viral mimicry induced by 5-aza-2-deoxycytidine. Proc Natl Acad Sci USA 113(37):10238–10244.
Rhie SK, Guo Y, Tak YG, Yao L, Shen H, Coetzee GA, Laird PW, Farnham PJ. 2016. Identification of activated enhancers and linked transcription factors in breast, prostate, and kidney tumors by tracing enhancer networks using epigenetic traits. Epigenetics Chromatin 9:50.
Zhang X, Guo C, Wu X, Li AX, Liu L, Tsark W, Dammann R, Shen H, Vonderfecht SL, Pfeifer GP. 2016. Analysis of liver tumor–prone mouse models of the Hippo kinase scaffold proteins RASSF1A and SAV1. Cancer Res 76(9):2824–2935.
2015
The Cancer Genome Atlas Research Network. 2015. The molecular taxonomy of primary prostate cancer. Cell 163(4):1011–1025.
Ciriello G, Gatza ML, Beck AH, Wilkerson MD, Rhie SK, Pastore A, Zhang H, McLellan M, Yau C, Kandoth C, Bowlby R, Shen H, Hayat S, Fieldhouse R, Lester SC, Tse GMK, Factor RE, Collins LC, Allison KH, Chen YY, Jensen K, Johnson NB, Oesterreich S, Mills GB, Cherniack AD, Robertson G, Benz C, Sander C, Laird PW, Hoadley KA, King TA, TCGA Research Network, Perou CM. 2015. Comprehensive molecular portraits of invasive lobular breast cancer. Cell 163(2):506–519.
Yao L, Shen H, Laird PW, Farnham PJ, Berman BP. 2015. Inferring regulatory element landscapes and transcription factor networks from cancer methylomes. Genome Biol 16:105.
Cancer Genome Atlas Research Network*. 2015. Comprehensive, integrative genomic analysis of diffuse lower-grade gliomas. New Engl J Med 372(26):2481–2498.
*Shen H is one of 328 co-authors
2014
Davis CF, Ricketts CJ, Wang M, Yang L, Cherniack AD, Shen H, Buhay C, Kang H, Kim SC, Fahey CC, Hacker KE, Bhanot G, Gordenin DA, Chu A, Gunaratne PH, Biehl M, Seth S, Kaipparettu BA, Bristow CA, Donehower LA, Wallen EM, Smith AB, Tickoo SK, Tamboli P, Reuter V, Schmidt LS, Hsieh JJ, Choueiri TK, Hakimi AA, The Cancer Genome Atlas Research Network, Chin L, Meyerson M, Kucherlapati R, Park WY, Robertson AG, Laird PW, Henske EP, Kwiatkowski DJ, Park PJ, Morgan M, Shuch B, Muzny D, Wheeler DA, Linehan WM, Gibbs RA, Rathmell WK, and Creighton CJ. 2014. The somatic genomic landscape of chromophobe renal cell carcinoma. Can Cell 26(3):319–330.
Hoadley KA, Yau C, Wolf DM, Cherniack AD, Tamborero D, Ng S, Leiserson MD, Niu B, McLellan MD, Uzunangelov V, Zhang J, Kandoth C, Akbani R, Shen H, Omberg L, Chu A, Margolin AA, Van’t Veer LJ, Lopez-Bigas N, Laird PW, Raphael BJ, Ding L, Robertson AG, Byers LA, Mills GB, Weinstein JN, Van Waes C, Chen Z, Collisson EA, The Cancer Genome Atlas Research Network, Benz CC, Perou CM, and Stuart JM. 2014. Multiplatform analysis of 12 cancer types reveals molecular classification within and across tissues of origin. Cell 158(4):929–944.
Cancer Genome Atlas Research Network.* 2014. Comprehensive molecular characterization of gastric adenocarcinoma. Nature 513(7517):202–209.
*Shen H is one of 315 co-authors
The Cancer Genome Atlas Research Network.* 2014. Comprehensive molecular characterization of urothelial carcinoma of the bladder. Nature 507(7492): 315–322.
*Shen H is a coauthor.
2013
Ari Hakimi A, Furberg H, Zabor EC, Jacobsen A, Schultz N, Ciriello G, Mikklineni N, Fiegoli B, Kim PH, Voss MH, Shen H, Laird PW, Sander C, Reuter VE, Motzer RJ, Hsieh JJ, Russo P. 2013. An epidemiologic and genomic investigation into the obesity paradox in renal cell carcinoma. J Natl Can Inst 105(24):1862–1870.
Yoshihara K, Shahmoradgoli M, Martnez E, Vegesna R, Kim H, Torres-Garcia W, Trevino V, The Cancer Genome Atlas Research Network, Shen H, Laird PW, Levine DA, Carter SL, Getz G, Stemke-Hale K, Mills GB, Verhaak RGW. 2013. Estimating the presence of tumor-associated normal cells using gene expression signatures predicts tumor purity. Nat Comm 4:2612.
Zack TI, Schumacher SE, Carter SL, Cherniack AD, Saksena G, Tabak B, Lawrence MS, Zhang CZ, Wala J, Hermel CH, Sougnez C, Gabriel SB, Hernandez B, Shen H, Laird PW, Getz G, Meyerson M, Beroukhim R. 2013. Pan-cancer patterns of somatic copy number alteration. Nat Gen 45(10):1134–1140.
Cancer Genome Atlas Research Network; Genome Characterization Center, …, Shen H, Triche T,…, Stuart JM. 2013. The Cancer Genome Atlas Pan-Cancer analysis project. Nat Gen 45(10):1113–1120.
Cheng WY, Ou Yang TH, Shen H, Laird PW, Anastassiou D, The Cancer Genome Atlas Research Network. 2013. Multi-cancer molecular signatures and their interrelationships. arXiv:1306.2584.
The Cancer Genome Atlas Research Network.* 2013. Comprehensive molecular characterization of clear cell renal cell carcinoma. Nature 499(7456):43–49.
*Shen H is a coauthor and led DNA methylation analysis.
The Cancer Genome Atlas Research Network.* 2013. Genomic and epigenomic landscapes of adult de novo acute myeloid leukemia. New Eng J Med 368(22):2059–2074.
*Shen H is a coauthor
The Cancer Genome Atlas Research Network, Kandoth C, Schultz N, Cherniack AD, Akbani R, Liu Y, Shen H, Robertson AG, Pashtan I, Shen R, Benz CC, Yau C, Laird PW, Ding L, Zhang W, Mills GB, Kucherlapati R, Mardis ER, Levine DA. 2013. Integrated genomic characterization of endometrial carcinoma. Nature 497(7447):67–73.
Shen H, Laird PW. 2013. Interplay between the cancer genome and epigenome. Cell 153(1):38–55.
Pharoah PD, …, Shen H,…, Sellers TA. 2013. GWAS meta-analysis and replication identifies three new susceptibility loci for ovarian cancer. Nat Gen 45(4):362–370.
Shen H,…, Laird PW, Goode EL, Leigh Pearce C. 2013. Epigenetic analysis leads to identification of HNF1B as a subtype-specific susceptibility gene for ovarian cancer. Nat Comm 4:1628.
2012
Lange CP, Campan M, Hinoue T, Schmitz RF, van der Meulen-de Jong AE, Slingerland H, Kok PJ, van Dijk CM, Weisenberger DJ, Shen H, Tollenaar RA, Laird PW. 2012. Genome-scale discovery of DNA-methylation biomarkers for blood-based detection of colorectal cancer. PLoS One 7(11):e50266.
The Cancer Genome Atlas Network.* 2012. Comprehensive molecular portraits of human breast tumors. Nature 490(7418):61–70.
*Shen H is a coauthor.
Zhang S, Liu CC, Li W, Shen H, Laird PW, Zhou XJ. 2012. Discovery of multi-dimensional modules by integrative analysis of cancer genomic data. Nuc Acids Res 40(19):9379–9391.
The Cancer Genome Atlas Network.* 2012. Comprehensive molecular characterization of human colon and rectal cancer. Nature 487(7407):330–337.
*Shen H is a coauthor.
Carter SL, Cibulskis K*, Helman E*, McKenna A*, Shen H*, Zack T*, Laird PW, Onofrio RC, Winckler W, Weir BA, Beroukhim R, Pellman D, Levine DA, Lander ES, Meyerson M, Getz G. 2012. Absolute quantification of somatic DNA alterations in human cancer. Nat Biotech 30(5):413–421.
*These authors contributed equally to the work.
Shen H, Laird PW. 2012. In epigenetic therapy, less is more. Cell Stem Cell 10(4):353–354.
2011
Campan M, Moffitt M, Houshdaran S, Shen H, Widschwendter M, Daxenbichler G, Long T, Marth C, Laird-Offringa IA, Press MF, Dubeau L, Siegmund KD, Wu AH, Groshen S, Chandavarkar U, Roman LD, Berchuck A, Pearce CL, Laird PW. 2011. Genome-scale screen for DNA methylation-based detection markers for ovarian cancer. PLoS One 29(12):1132–1144.
International Stem Cell Initiative,…, Laird PW, …, Shen H, …, Zhou Q. 2011. Screening ethnically diverse human embryonic stem cells identifies a chromosome 20 minimal amplicon conferring growth advantage. Nat Biotech 29(12):1132–1144.
Hinoue T, Weisenberger DJ, Lange CP, Shen H, Byun HM, Van Den Berg D, Malik S, Pan F, Noushmehr H, van Dijk CM, Tollenaar RA, Laird PW. 2012. Genome-scale analysis of aberrant DNA methylation in colorectal cancer. Gen Res 22(2):271–282.
The Cancer Genome Atlas Research Network. 2011. Integrated genomic analyses of ovarian carcinoma. Nature 474(7353):609–615.
*Shen H is a coauthor and led the DNA methylation analysis.
Oghamian S, Sodir NM, Bashir MU, Shen H, Cullins AE, Carroll CA, Kundu P, Shibata D, Laird PW. 2011. Reduction of pancreatic acinar cell tumor multiplicity in Dnmt1 hypomorphic mice. Carcinogen 32(6):829–835.
Noushmehr H,Weisenberger DJ, Diefes K, Phillips HS, Pujara K, Berman BP, Pan F, Pelloski CE, Sulman EP, Bhat KP, Verhaak RG, Hoadley KA, Hayes DN, Perou CM, Schmidt HK, Ding L, Wilson RK, Van Den Berg D, Shen H, Bengtsson H, Neuvial P, Cope LM, Buckley J, Herman JG, Baylin SB, Laird PW, Aldape K; Cancer Genome Atlas Research Network. 2010. Identification of a CpG island methylator phenotype that defines a distinct subgroup of glioma. Can Cell 17(5):510–522.
Teschendorff AE, Menon U, Gentry-Maharaj A, Ramus SJ, Weisenberger DJ, Shen H, Campan M, Noushmehr H, Bell CG, Maxwell AP, Savage DA, Mueller-Holzner E, Marth C, Kocjan G, Gayther SA, Jones A, Beck S, Wagner W, Laird PW, Jacobs IJ, Widschwendter M. 2010. Age-dependent DNA methylation of genes that are suppressed in stem cells is a hallmark of cancer. Gen Res 20(4):440–446.
DNA Methylation Analysis Tools
BISCUIT
The BISulfite-seq CUI Toolkit (BISCUIT) is a tool for analyzing sodium bisulfite conversion-based DNA methylation/modification data. It was written to align, perform DNA methylation extraction and call single nucleotide mutations from cytosine-converted sequencing data.
Access BISCUIT ➔
Related Publication
Zhou W, Johnson BK, Morrison J, Beddows I, Eapen J, Katsman E, Semwal A, Habib W, Heo L, Laird PW, Berman B, Triche Jr TJ, Shen H. 2024. BISCUIT: an efficient, standards-compliant tool suite for simultaneous genetic and epigenetic inference in bulk and single-cell studies. Nuc Acids Res 52(6):e32.
BISCUIT Snakemake Workflow
The BISCUIT Snakemake Workflow is a Snakemake-based pipeline to run BISCUIT alignment and DNA methylation extraction across all samples in an experiment. It also provides many optional steps including a variety of quality control steps.
Access BISCUIT Snakemake Workflow ➔
Dupsifter
Dupsifter is a command line tool for marking PCR duplicates in both WGBS and WGS datasets.
Access Dupsifter ➔
Related Publication
Morrison J, Zhou W, Johnson BK, Shen H. 2023. Dupsifter: A lightweight duplicate marking tool for whole genome bisulfite sequencing. Bioinformatics 39(12):btad729.
SeSAMe
SeSAMe is an R/Bioconductor package for processing Illumina Infinium DNA methylation array data.
Access SeSAMe on GitHub ➔
Access SeSAMe on Bioconductor ➔
Related Publication
Zhou W, Triche Jr. TJ, Laird PW, Shen H. 2018. SeSAMe: reducing artifactual detection of DNA methylation by Infinium BeadChips in genomic deletions. Nuc Acids Res 46(20):e123-e123.
RNA-Sequencing Tools
STORMqc
STORMqc is a Snakemake-based pipeline for quality control of STORM-seq data such as finding reads mapped to undesirable locations, basic read quality metrics and number of reads.
Access STORMqc ➔
Tranquillyzer
Tranquillyzer (TRANscript QUantification In Long reads-anaLYZER) is a flexible, architecture-aware deep learning framework for processing long-read single-cell RNA-seq data.
Access Tranquillyzer ➔
Tranquillyzer-nf
Tranquillyzer-nf is a Nextflow-based pipeline for reproducible processing of long-read single-cell RNA-seq experiments with Tranquillyzer.
Access Tranquillyzer-nf ➔
General Tools
iscream
iscream is an R/C++ package for efficiently reading BED files into formats usable within the R/Bioconductor framework. It is optimized to query genomic regions, summarize the queried data and make matrices.
Access iscream on GitHub ➔
Access iscream on Bioconductor ➔
Synthbar
Synthbar adds synthetic sequences such as cell barcodes or unique molecular indexes to reads to make them conform to an expected format for downstream analysis.
Access Synthbar ➔
Related Publication
Morrison J, Johnson BK, Shen H. 2025. Synthbar: A lightweight tool for adding synthetic barcodes to sequencing reads. BioRxiv.


Ian Beddows, Ph.D.
Senior Bioinformatics Research Scientist, Bioinformatics and Biostatistics Core

Svetlana Djirackor
Ph.D. Student, VAI Graduate School
Research Focus: Parallel profiling of the epigenomic and transcriptomic landscape across the three major ovarian cancer histotypes

James Eapen
Ph.D. Student, VAI Graduate School
Research Focus: Developing tools for genomic data analysis

Kelly Krzyzanowski, B.S.
Laboratory Manager, Department of Epigenetics

Ben Johnson, Ph.D.
Senior Research Scientist, Department of Epigenetics & Department of Cell Biology
Epigenetics of reproductive cancers

Paige Matusiak
Ph.D. Student, VAI Graduate School
Research Focus: Investigating transposable element activity in ovarian cancer subtypes

Jacob Morrison, Ph.D.
Bioinformatics Research Scientist, Department of Epigenetics

Amy Nuffesse
Senior Administrative Assistant II, Department of Epigenetics


Mary Rhodes, B.S.
Senior Lab Manager, Department of Epigenetics

Ayush Semwal, M.S.
Bioinformatics Analyst II, Department of Epigenetics

Natasha Zandstra
Ph.D. Student, VAI Graduate School
Research Focus: Identifying non-genetic determinants of stem-like phenotypes in high grade serous ovarian cancer


